For our international customers, please be advised that orders cannot be placed through our website by customers in countries with International Distributor representation.
Amylase, Alpha - Manual
Porcine pancreatic alpha-amylase (PPA) acts upon large linear carbohydrate polymers at internal bonds. The hydrolytic products have alpha-configuration. alpha-Amylase activity is present in all living organisms; however, the enzymes vary remarkably even from tissue to tissue within a single species.
The alpha-amylases were named by Kuhn in 1925, because the hydrolysis products are in the alpha configuration. In 1930, Ohlsson discovered another amylase, which yielded a beta-mannose. He named it beta-amylase. In 1947, the enzyme was first crystallized from pig pancreas by Meyer et al., and Danielsson determined its molecular weight. In 1952, Caldwell et al. reported on an improved purification method.
Inhibitors of alpha-amylase were reported on as early as 1975 (Marshall and Lauda 1975). The first crystal structure of alpha-amylase was from Aspergillus oryzae and was determined in 1979 (Matsuura et al. 1979). This 6 Å resolution structure was improved to a 3 Å resolution the following year (Matsuura et al. 1980).
In the 1980s, the amino acid sequences of the two forms (PPAI and PPAII) were determined (Kluh et al. 1981, and Pasero et al. 1986). The three dimensional crystal structures of each form were determined in the 1990s and found to be effectively identical (Qian et al. 1993, and Gilles et al. 1996). The complete cDNA sequence was determined by Darnis et al. in 1999.
Recent work with alpha-amylase has included isolating and studying the effects of inhibitors from natural sources (Narita and Inouye 2009, and Suda et al. 2011). Researchers have also studied the effects of bound carbohydrates on enzyme stability (Gopal et al. 2008) and analyzed the binding of inhibitors with the aim of designing novel inhibitors (Najafian et al. 2011).
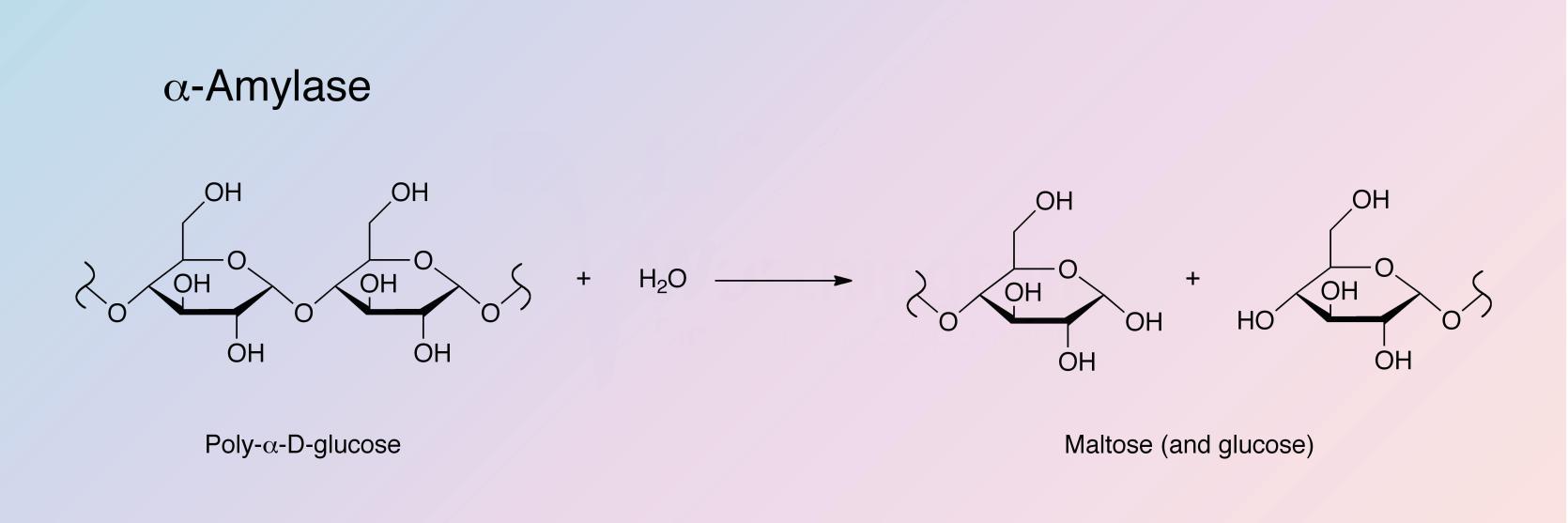
alpha-Amylase catalyzes the hydrolysis of internal alpha-1,4-glucan links in polysaccharides containing 3 or more alpha-1,4-linked D-glucose units, yielding a mixture of maltose and glucose. See also Takeshita and Hehre (1975).
The enzyme is a glycoprotein (Beaupoil-Abadie et al. 1973). Its single polypeptide chain of about 475 residues has two free thiol groups, four disulfide bridges, and contains a tightly bound Ca2+ (Granger et al. 1975, and Steer et al. 1974). It exists in two forms (PPAI and PPAII), which have identical enzymatic properties and molecular mass of 55.4 kDa, differing only in electrophoretic mobility and pI (Cozzone et al. 1970b, Marchis-Mouren and Pasero 1967, and Darnis et al. 1999).
The structures of both forms of PPA consist of a major domain A (residues 1-99 and 170-404) organized as an (alpha/beta)8-barrel that contains a loop (domain B). Following the C-terminal domain C (residues 405-496) is a ten-beta-stranded Greek key motif (Darnis et al. 1999).
Animal alpha-amylases are encoded by two loci, amy1 (salivary) and amy2 (pancreatic). The full length amy cDNA contains 1565 bp. It contains a 15 amino acid signal peptide and a 1536-nt open reading frame (ORF). A 29-nt 3’ non-coding region is followed by a short poly(A) tail.
Animal alpha-amylases contain five conserved regions including four at or around the (alpha/beta)8-barrel and a short sequence near the C-terminus of domain B (Janecek 1993, Svensson 1994, and Janecek and Balaz 1995). Human and porcine pancreatic alpha-amylases are 87.1% identical, with only 64 amino acid substitutions. Mouse and rat alpha-amylase share a 85.5% identity with that of porcine. Amino acids 304-310 are conserved among mammalian alpha-amylases and are believed to be involved in the enzymatic mechanism (Darnis et al. 1999).
- Ethanol production
- Reduction of starch or glycogen molecules
P00690
- Class: Alpha Beta; Mainly Alpha
- Architecture: Alpha-Beta Barrel; Sandwich
- Topology: TIM Barrel; Immunoglobulin-like
- 51.0-54.0 kDa (Cozzone et al. 1970a)
- 55.4 kDa (SDS-PAGE) (Alkazaz et al. 1996)
7.0
- PPAI: 7.5 (Ajandouz et al. 1995)
- PPAII: 6.4 (Ajandouz et al. 1995)
- 133,870 cm-1 M-1 (Theoretical)
- E1%, 280 = 26 (Caldwell et al. 1952)
- Chloride is essential. (See also O’Donnell et al. 1975 for the effect of bile salts)
- Calcium and chloride ions are necessary for stability.
- Phenolic compounds (Funke and Melzig 2005)
- Plant extracts (Karthic et al. 2008, and Buonocore et al. 1985)
- Urea and other amide reagents (Toralballa and Eitingon 1967)