For our international customers, please be advised that orders cannot be placed through our website by customers in countries with International Distributor representation.
Beta Agarase - Manual
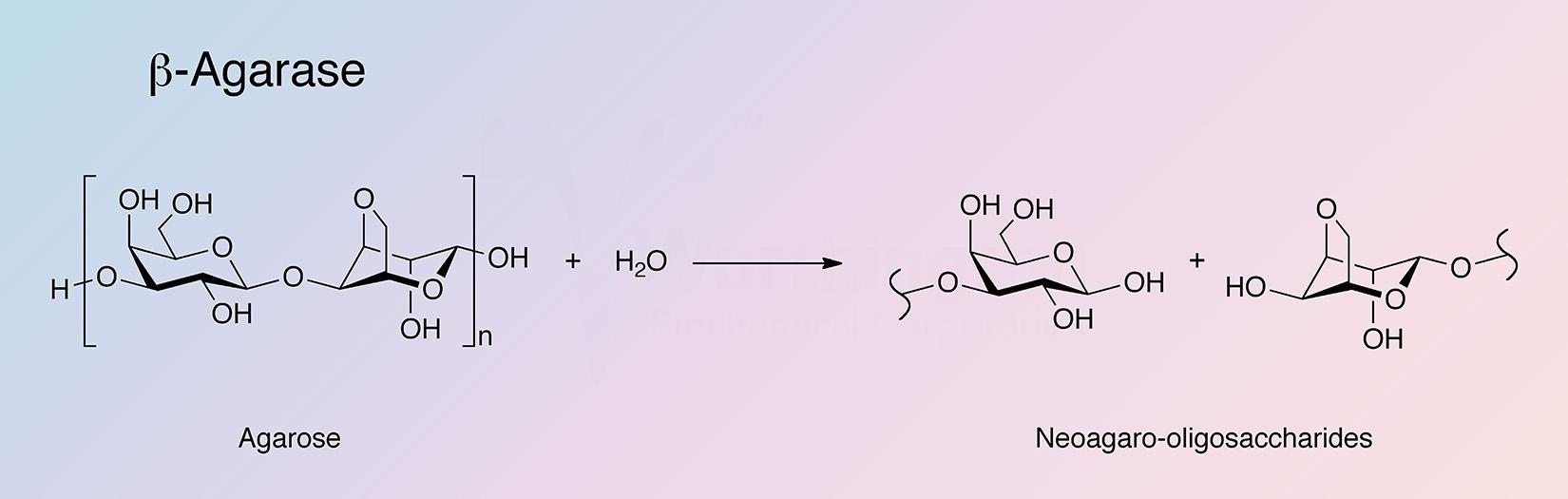
Agarases are glycoside hydrolases that catalyze the hydrolysis of agar. Agarases are primarily found in marine organisms (Fu and Kim 2010), with microorganisms being the primary sources for agarase production (Oh et al. 2010). Agarases are classified into two groups based on their mode of action. α-Agarases hydrolyze α-1,3-linkages to produce agarooligosaccharides of a series related to agarobiose, while β-agarases hydrolize β-1,4-linkages producing neoagarooligosaccharides of a series related to neoagarobiose (Fu and Kim 2010, and Oh et al. 2010).
Yaphe and colleagues first described the agarose-degrading system of Pseudomonas atlantica (Yaphe 1957, Yaphe 1966, Day and Yaphe 1975, and Groleau and Yaphe 1977). This enzyme was designated β-Agarase II. The purification and characterization of the related β-Agarase I (32 kDa) was reported by Morrice et al. in 1983. The two enzymes differ significantly in that β-Agarase I is membrane-bound, while β-Agarase II is found extracellularly (Morrice et al. 1983a and b). That formerly manufactured by Worthington Biochemical was β-agarase II.
Belas et al. cloned β-agarase II from Pseudomonas atlantica gene in 1988, and the nucleotide sequence was determined in 1989 (Belas et al. 1988, and Belas et al. 1989). Worthington Biochemical first manufactured β-agarase II in the early 1990s.
Recent research has involved the identification of novel β-agarase sequences from agar-degrading bacterial genomes (Lee et al. 2015a and b) and the characterization of newly discovered agarases (Chi et al. 2015, and Hsu et al. 2015). In addition, the crystal structures of β-agarase enzymes continue to be elucidated (Takagi et al. 2014, and Pluvinage et al. 2013).
β-Agarase is unique in its ability to break down the agarose polysaccharide core composed of repeating 1,3-linked β-D-galactopyranose and 1,4-linked 3,6-anhydro-alpha-L-galactopyranose into neoagarobiose oligosaccharides.
Based on catalytic domain similarities, β-Agarase II of Pseudoalteromonas atlantica has been designated part of the glycosyl hydrolase 86 (GH86) family. Other β-agarases belong to glycosyl hydrolase families 16 and 50, with all three groups containing a conserved glycoside hydrolase region responsible for catalysis (Fu and Kim 2010).
The NH2 terminus of the protein may function as a signal peptide in the export of extracellular β-agarase. Signal peptidase is believed to cleave between resides 22 and 23 of the primary agrA product, yielding a mature protein of 482 residues (Belas et al. 1989).
The gene encoding β-agarase (agrA) has a primary structure consisting of an ORF of 1,515 nucleotides. The primary translation product consists of 504 amino acids (Belas 1989). Three regions upstream to the start codon that are responsible for ribosomal binding and promoter sites of the gene sequence have been determined based on homology with E. coli (Belas et al. 1989).
- Food, cosmetic and medical industry (Kobayashi et al. 1997)
- Protoplast isolation from seaweeds (Araki et al. 1998)
- Production of agar-derived oligosaccharides
- DNA recovery from agarose gel: As a result of this enzymatic degradation the viscosity and gelling ability of the native low melting point agarose is eliminated liberating the nucleic acids. Since the oligosaccharides produced are alcohol soluble, intact DNA can be recovered in a simple and efficient manner by alcohol precipitation.
P13734
- Class: Mainly beta
- Architecture: Sandwich
- Topology: Jelly Rolls
- 55.1 kDa (Theoretical, mature chain)
6.5-7.5 (Morrice et al. 1983b)
- 5.9 (Oh et al. 2010)
- 5.95 (Theoretical)
- 97,250 cm-1 M-1 (Theoretical)
- E1%,280 = 17.66 (Theoretical)
- Glutamic Acid (E177 and E299)