For our international customers, please be advised that orders cannot be placed through our website by customers in countries with International Distributor representation.
Aldolase - Manual
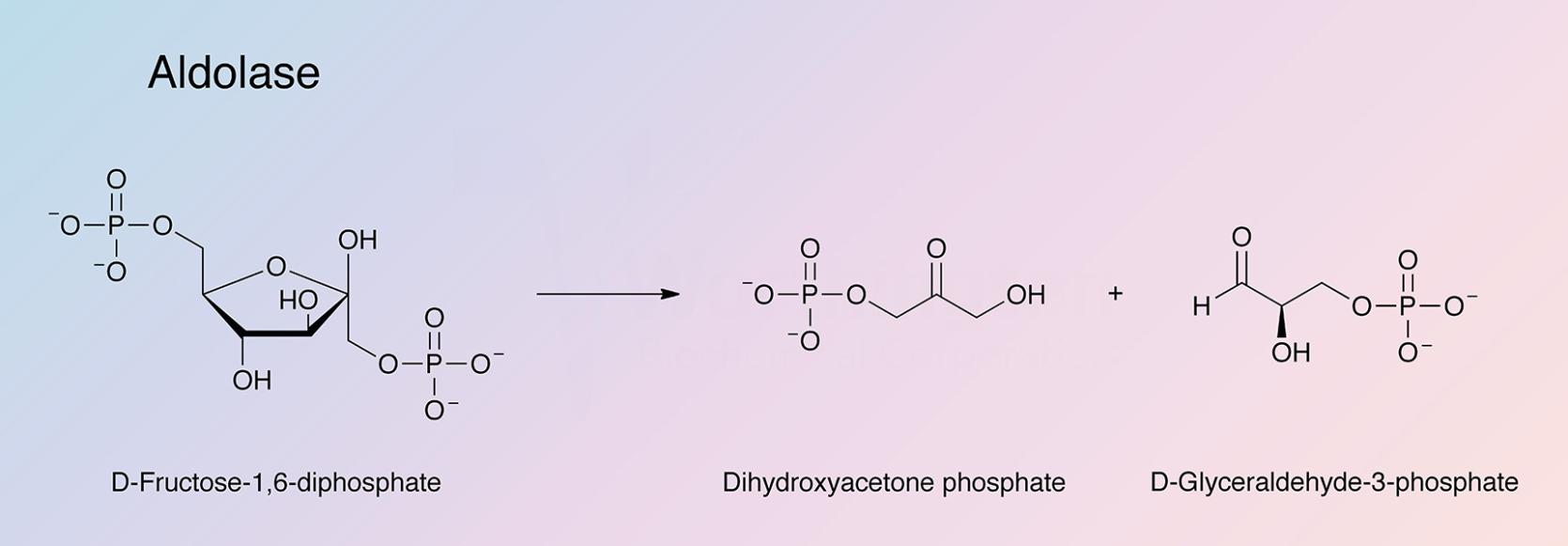
Fructose-biphosphate aldolase (ALD) catalyzes a key reaction in glycolysis and energy production:
Aldolase is present in all animal and plant tissue, and in most microorganisms. Class I aldolase, found in animal and higher plant tissue, is characterized by the lack of requirement for a bivalent metal cofactor, and by the formation of ketimine Schiff base intermediate with the substrate dihyroxyacetone phosphate. Class II aldolase, found only in prokaryotes and lower eukaryotes, requires a bivalent metal cofactor (Heron and Caprioli 1975, London 1974, and Lebherz et al. 1973, Gefflaut et al. 1995). There are three types of Class I aldolases: Type A (the major form) is found in muscle; Type B in liver and kidney; and Type C (plus some A) in brain (Horecker et al. 1972).
In 1934, aldolase was found to be present in muscle and yeast by Meyerhof and Lohmann. Muscle aldolase was first separated from isomerase by Herbert et al. in 1940. Neuberg and Kobel first proposed the presence of aldolase in bacteria in 1934, which was later confirmed in E. coli (Utter and Werkman 1941).
In the 1950s and 1960s the reaction mechanisms were investigated, and the classification scheme that divided aldolases into Class I and Class II was developed (Marsh and Lebherz 1992).
Work investigating the inhibitors began in the mid-1960s (Spolter et al. 1965), and early 1970s (Lehrer and Barker 1970, and Hinz et al. 1971). In 1969, the crystals of aldolase were studied in order to determine the tetrameric subunit structure (Eagles et al. 1969). The first low resolution crystal structure was determined in 1985 (Sygusch et al. 1985), and was refined in 1987 (Sygusch et al. 1987).
After some debate over the mechanism and the involvement of particular active side residues, Littlechild and Watson determined in 1993 that a cysteine residue was not present at or near the substrate binding site, despite initial studies concluding otherwise (Littlechild and Watson 1993). The first high resolution structure of rabbit aldolase was reported in 1997 (Blom and Sygusch 1997).
Today, work investigating aldolase inhibitors continues (Dax et al. 2005). Studies are also being conducted to better understand its utility in chemo-enzymatic syntheses (Bolt et al. 2008, and Calveras et al. 2009), its interactions with other cellular proteins (Forlemu et al. 2007), and its reaction intermediates (St.-Jean et al. 2005).
Aldolase is highly specific for its substrate, D-fructose.
The three isozymes (A, B, and C) found in mammals are composed of four identical subunits, each a polypeptide chain of 360 amino acids. The enzyme has a homotetrameric structure with subunits that are approximately 40 kDa (Marsh and Lebherz 1992). The structure of these subunits is a beta-barrel, surrounded by alpha-helices and connecting loops (Littlechild and Watson 1993). If activity of one of the four subunits of the homotetramer is knocked out, the entire complex is incapable of normal function, indicating a tight communication between active sites (Marsh and Lebherz 1992).
The primary structures of Class I eukaryotic enzymes are highly conserved (Kelly and Tolan 1986, and Freemont et al. 1988); however, there are no similarities between the sequences of Class I and Class II enzymes, as they have separate evolutionary origins (Marsh and Lebherz 1992). However, both Class I and Class II enzymes contain carboxy-terminal tyrosine residues that are required for maximal catalytic activity (Rutter 1961, and Morse and Horecker 1968). Prolonged contractile activity reduces the type A aldolase expression in mRNA by downregulating cytoplasmic glycolytic enzymes, and enhancing mitochondrial enzymatic activity (Williams et al. 1986).
- Chemo-enzymatic syntheses (Bolt et al. 2008 and Calveras et al. 2009)
- Structure-function studies (St.-Jean et al. 2005)
- Organic synthesis (Castillo et al. 2006, Concia et al. 2009)
- Quantification of D-fructose-1,6-diphosphate
- Metabolite determination in coupled reactions (Hiller et al. 2007)
- Molecular weight marking in columns and gels
P00883
- Class: Alpha Beta
- Architecture: Alpha-Beta Barrel
- Topology: TIM Barrel
- 156.8 kDa (Theoretical)
6.9-8.8
8.47 (Theoretical)
- 133,560 cm-1 M-1 (Theoretical)
- E1%,280 = 8.52 (Theoretical)
- Lysine (K229)
- Glutamate (E187)
- Plus other charged residues (functional assignment is complex due to the close proximity of the many charged amino acids found at the active site) (St.-Jean et al. 2005)
- Heavy metals, especially Cu2+, Zn2+, and Ag+
- Hydroxynaphthaldehyde phosphate derivatives (Dax et al. 2005)
- Phosphorylated aromatic compounds (Blonski et al. 1997)
- D-Erythrulose 1-phosphate (Ferroni et al. 1991)
- Phosphatidylserine liposomes (Kwiatkowska et al. 1994)
- Phosphates and adenine nucleotides (Callens et al. 1991)