For our international customers, please be advised that orders cannot be placed through our website by customers in countries with International Distributor representation.
Asparaginase - Manual
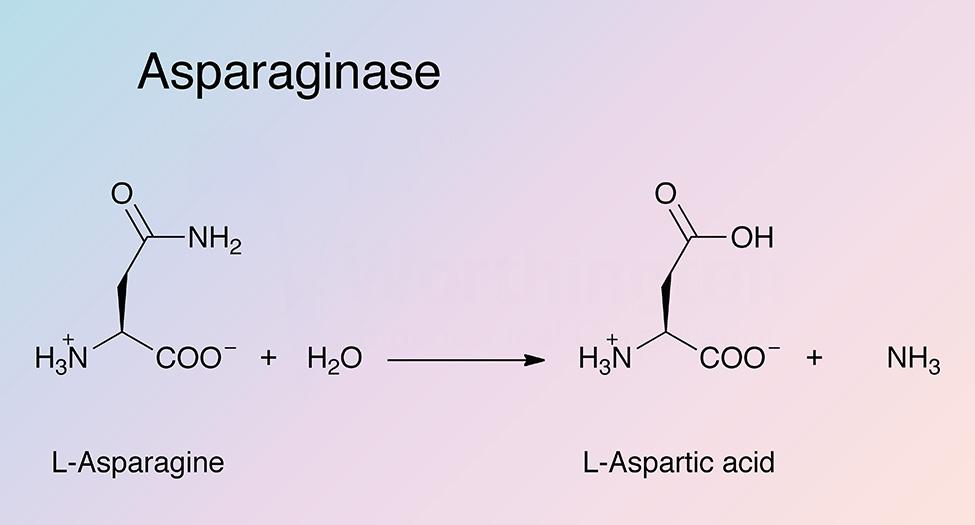
Asparaginase catalyzes the hydrolysis of L-asparagine to produce L-aspartic acid and ammonia. Bacterial type L-asparaginases are classified into subtypes I and II, which is defined by their intra- or extracellular localization (Michalska and Jaskolski 2006). Type I (cytosolic) has a lower affinity for L-asparagine, while type II (periplasmic) has a high substrate affinity. Plant type L-asparaginases differ structurally and have a different evolutionary origin than bacterial L-asparaginases (Michalska et al. 2006)
In 1904, Lang became the first to detect asparaginase activity in beef tissues. Fürth and Friedmann confirmed these results in 1910. Fürth and Friedmann believed asparaginase activity was found at the same level in all animal tissues, which was later refuted by Clementi (Clementi 1922) whose experiments concluded the enzyme was active in practically all tissues of herbivores, only in the liver of omnivorous animals, and absent in the organs of carnivorous animals, amphibians, and reptiles (Michalska and Jaskolski 2006).
In 1953, Kidd reported anti-tumor properties of guinea pig serum, which was later attributed to asparaginase activity (Broome 1961). In 1963, Mashburn and Wriston found that the E. coli enzyme had anti-tumor activity. Work proceeded in several laboratories leading to clinical trials, making E. coli asparaginase II the first antileukemic L-asparaginase to be used clinically (Roberts et al. 1966, Old et al. 1967, Boyse et al. 1967, Hill et al. 1967, Clarkson et al. 1970, and Oettgen et al. 1970). It was soon discovered that in addition to the secreted EcAII, E. coli (and other bacteria) also produce cytosolic asparaginase, asparaginase I (EcAI), which has a lower affinity for L-asparagine and is not effective against cancer (Willis and Woolfolk 1974).
Preliminary efforts to determine the primary structure were carried out in 1974, and the sequence was confirmed in 1990 (Maita et al. 1974, Maita and Matsuda 1980, and Bonthron 1990). After crystallographic studies by Epp et al. in 1971, a partial model of the EcAII crystal structure was published in 1988 by Ammon et al.. Although not completely accurate, this structure was used as a stepping stone by Swain et al. who elucidated the true structure in 1993. Many crystal structures of L-asparaginase from a wide variety of organisms have been determined (Lubkowski et al. 2003).
Current research with L-asparaginase continues to focus on its oncological importance, and the specific transcription factors EcAII targets (Suto et al. 2010). Structure-function relationships of L-asparaginase from a variety of sources are also currently under investigation (Labrou and Papageorgiou 2010, and Bansal et al. 2010).
EcAII exhibits a broad pH activity profile and higher substrate affinity than EcAI. The reaction consists of two steps. In the first step, the nucleophilic threonine attacks the carbonyl of the amide substrate to generate an acyl-enzyme intermediate. A molecule of ammonia is released. In the second step, water attacks the acyl-enzyme intermediate to produce L-aspartate (Michalska and Jaskolski 2006).
EcAII is a tetrameric protein composed of four identical subunits, each subunit contains 326 amino acid residues. The two threonine residues present at the active site are required for activity (see PDB 3ECA). Debate still continues over which threonine residue supplies the nucleophile. Most believe that it is the N-terminal threonine. The C-terminal threonine is part of a triad that resembles the typical SHD catalytic triad of serine proteases (Carter and Wells 1988).
The promoter of the gene encoding L-asparaginase II, ansB, is coactivated by cAMP receptor protein (CRP) and the FNR protein (Scott et al. 1995). CRP is activated in response to carbon sources, and the FNR protein, the fnr gene product, in response to anaerobiosis. The FNR site is located 41.5 bp upstream of the major start site, and 28 bp upstream of the FNR site is the CRP regulation site (Jennings and Beacham 1993).
- Treatment of acute lymphoplastic leukemia (Zhang et al. 2004)
- L-asparagine concentration determination (Gentili et al. 1994)
P00805
- Class: Alpha Beta
- Architecture: 3-Layer(aba) Sandwich
- Topology: Rossmann Fold
- 138.4 (Theoretical)
- 8.6 (Zhang et al. 2004)
- 8.0 (Wriston 1971)
- 5.67 (Theoretical)
- 85,160 cm-1 M-1
- E1%, 278 = 7.1 (Ho et al. 1971)
- Threonine (T34, T111)
- Aspartate (D112)
- Lysine (K184)
- Anaerobic conditions (Cedar and Schwartz 1967)
- Amino acids (not specifically asparagine) (Cedar and Schwartz 1968)
- 5-diazo-4-oco-L-norvaline (DONV) (Wriston and Yellin 1973)
- Dimethylsulfoxide (Wriston 1971)
- Glutaraldehyde (Zhang et al. 2004)
- L-Asparagine and L-aspartate (Wriston and Yellin 1973)
- NH4+ (Wriston 1971)
- Trypsin (Zhang et al. 2004)