For our international customers, please be advised that orders cannot be placed through our website by customers in countries with International Distributor representation.
Amylase, Beta - Manual
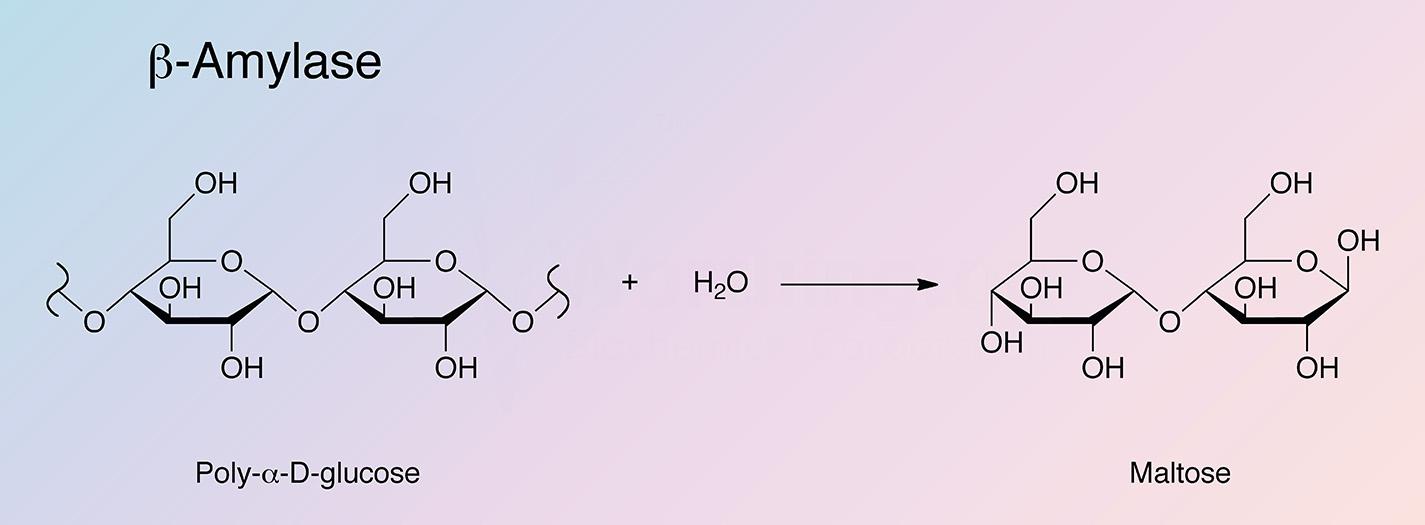
β-Amylase is an exoenzyme that releases successive maltose units from the nonreducing end of a polysaccharide chain by hydrolysis of α-1,4-glucan linkages. The shortest normal saccharide attacked is maltotetraose (Myrback and Neumuller 1950). Since it is unable to bypass branch linkages in branched polysaccharides such as glycogen or amylopectin, the hydrolysis is incomplete and a macromolecular limit dextrin remains.
β-Amylase is found primarily in the seeds of higher plants and sweet potatoes. It yields a single product: maltose. Marshal and Whelan (1973) report on the removal of any contaminating β-glucosidase. The tuberous root of sweet potato is unusually rich in the enzyme, accounting for approximately 5% of the total soluble proteins. In contrast, other tuberous roots only contain trace amounts of β-amylase activity (Li and Oba 1985, and Yoshida and Nakamura 1991).
The α-amylases were named by Kuhn in 1925, because the hydrolysis products are in the alpha configuration. In 1930, Ohlsson discovered another amylase, which yielded a β-mannose and named it β-amylase.
β-amylase was first purified in the 1940s (Schoch 1942, and Haworth et al. 1946). Hopkins et al. first began to study the kinetics of β-amylase in 1948.
In the 1960s and 1970s, techniques in the purification of sweet potato β-amylase improved (Nakayama and Amagase 1963, Takeda and Hizukuri 1969, and Hegde et al. 1979). The enzyme was successfully immobilized on agarose and sepharose gels in the late 1970s (Caldwell et al. 1976a, b).
In 1991, β-amylase was successfully cloned and expressed in E. coli (Yoshida et al. 1991). The complete amino acid sequence and preliminary crystal structure of the tetrameric enzyme were determined in 1993 (Toda et al. 1993, and Cudney and McPherson 1993). The crystal structure was refined in 1995 (Cheong et al. 1995). In 1996, Pujadas et al. studied the evolution of β-amylases determining the enzymes to be an example of parsimonious divergence illustrated by signature structural motifs.
In 2001, a simple purification method using affinity precipitation was developed (Teotia et al. 2001). The effects of guanidine hydrochloride and increased pressure on the activity of the enzyme were studied in 2002 (Tanaka et al. 2002).
Most β-amylases are monomeric enzymes (Thoma et al. 1971). However, that of sweet potato is tetrameric consisting of four identical subunits (Thoma et al. 1963). Each subunit consists of a large (α/β)8-barrel region also seen in a-amylase. A smaller, globular region is formed by long loops extending from β-strands. Between these two regions is a cleft believed to open and contain the conserved Glu187. Cys96 is located at the entrance of the (α/β)8-barrel region and is important to inactivation by sulfhydryl reagents (Cheong et al. 1995).
The gene that encodes for β-amylase is amyB. A 1404 bp open reading frame encodes for the 499 amino acid precursor to the β-amylase subunit. After methionine processing, the mature form of the protein contains 498 residues (Yoshida and Nakamura 1991). The gene contains seven exons and six introns, and a TATA box sequence is present 26 to 30 bp upstream of the multiple transcription sites. The upstream region of amyB contains sequences to which several known plant nuclear factors bind; regulation of the amyB gene is thought to be controlled by one or more of the common cis-regulatory elements (Yoshida et al. 1992). Subunits of sweet potato β-amylase and seed β-amylase from barley and soybean share approximately 68% amino acid identity (Yoshida and Nakamura 1991).
- Starch and glycogen structural studies
- Fermentation in brewing and distilling industry
- Saccharification of liquefied starch: production of high maltose syrups and high conversion syrups
P10537
- Class: Alpha Beta
- Architecture: Alpha-Beta Barrel
- Topology: TIM Barrel
- 223.8 kDa (Theoretical)
- 4.0-5.0 (French 1960)
- 5.5 (Ohba et al. 1979)
- 5.17 (Theoretical)
- 388,640 cm-1 M-1 (Theoretical)
- E1%, 280 = 17.7 (Takeda and Hizukuri 1969)
- Glutamate (G187 and G382)
- Heavy metal ions
- PCMB and PMSF
- Iodoacetamide
- Ascorbate (Rowe and Weill 1962)
- Cyclohexaamylose
- Sulfhydryl reagents