For our international customers, please be advised that orders cannot be placed through our website by customers in countries with International Distributor representation.
Carboxypeptidase B - Manual
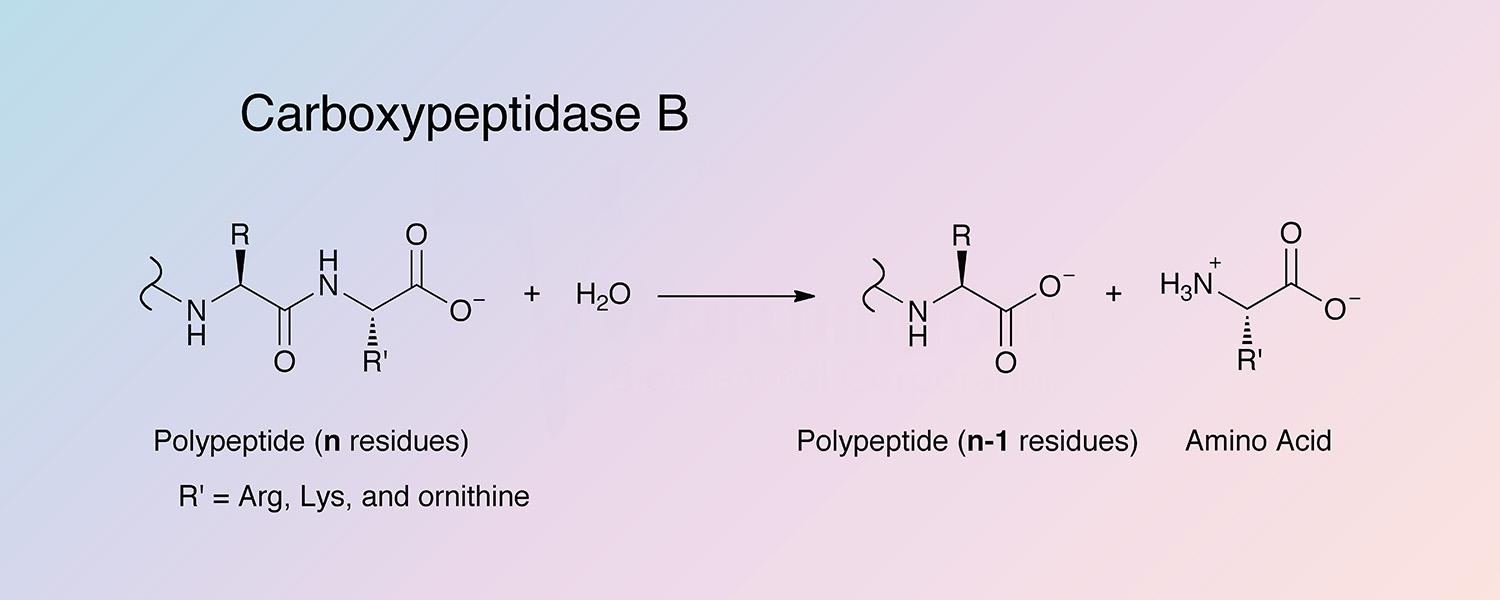
Carboxypeptidase B (CPDB) is a metallocarboxypeptidase that catalyzes the hydrolysis of the basic amino acids, lysine, arginine, and ornithine from the C-terminal position of polypeptides:
Peptidyl-L-lysine(-L-basic amino acid) + H2O --> Peptide + L-lysine(L-basic amino acid)
Carboxypeptidases are secreted as zymogens by the acinar cells of the pancreas. The zymogens are activated by trypsin in the small intestine..
In 1931, Waldschmidt-Leitz et al. demonstrated that excretions from porcine pancreas contained an enzyme that catalyzed the release of arginine from proteins (Folk 1971). They termed the enzyme “protaminase”. In 1933, Calvery used egg albumin to show that it too could be susceptible to this enzyme, suggesting that lysine and histidine could also be released. Throughout the 1930s and 1940s, it was believed that this enzyme was identical to chymotrypsin (Sumner and Somers 1947, and Northrop et al. 1948).
In 1940, the work of Hoffmann and Bergmann demonstrated lysine was only slowly released from substrate in the presence of carboxypeptidase A (CPDA). The work of Folk in 1956 showed lysine was rapidly released in the presence of commercially available pancreas powder (Folk 1971). These two insights led to experiments that confirmed the presence of two separate carboxypeptidase enzymes (Folk 1956, and Folk and Gladner 1958). The carboxypeptidase B name was given by Folk et al. in 1960 in order to officially distinguish it from CPDA.
In 1975, Titani et al. determined the amino acid sequence. In 1976, Schmid and Herriott resolved the structure to a 2.8 Å resolution, which was later refined by Coll et al. in 1991. The globular domain of pancreatic porcine pro-CPDB pro-segment was derived through NMR spectroscopy by Vendrell et al. in 1990.
Recent work has included obtaining a higher resolution crystal structure of the proform of CPDB (Fernández et al. 2010), as well as resolving the crystal structure of potent thiol inhibitors (Adler et al. 2005).
CPDB is activated by trypsin. CPDB is highly specific for lysine and arginine, but shows preference for arginine (Tan and Eaton 1995). It can also act (at a slower rate) on valine, leucine, isoleucine, asparagine, glycine, and glutamine (Villegas et al. 1995, Nishihira et al. 1995). The differences in specificities between carboxypeptidase A (CPDA) and CPDB can be attributed to the residues Ser205, Gly241, and Asp253 in CPDB as compared to Gly207, Ile243, and Ile255 in CPDA (Coll et al. 1991).
The major form of CPDB is found in the monomeric state. CPDB contains 1 atom of zinc per mole of protein. The residues coordinating the zinc residue are conserved and include two histidines, a glutamic acid, and a water (Avilés and Vendrell 2004).
Clauser et al. studied the rat preprocarboxypeptidase B gene, and found that it is organized very similarly to the bovine CPB1 gene (Clauser et al.1988). The CPB1 gene is conserved in many eukaryotes and has been studied in human, chimpanzee, dog, mouse, rat, chicken, zebrafish, and fruit fly (Avilés and Vendrell 2004). The mature form of rat CPDB is homologous to bovine CPDB (77% identical), and the amino acids involved in catalysis or ligand binding show no variance. The coding region of preprocarboxypeptidase B consists of 11 exons (Clauser et al. 1988). Comparisons of rat CPDB, rat CPDA, and rat CPDA2 genes have shown that the number, position, and sequence composition of the exons are conserved, despite large differences in intervening sequences (Gardell et al. 1988, and Clauser et al. 1988). Through cDNA and mRNA sequencing, CPDB mRNA has been determined to be 1385 nucleotides, excluding the poly(A) tail. The rat CPDB mRNA is 100 nucleotides longer the CPDA mRNA. This difference is primarily due to a larger 3’ untranslated region (UTR) in the CPDB mRNA (Han et al. 1986).
- Sequence analysis by successive cleavage of C-terminal basic amino acids
- Serum marker for acute pancreatitis diagnosis (Avilés and Vendrell 2004)
- Cytochrome C stepwise digestion (Südi et al. 1989)
- Human insulin production (Ladisch and Kohlmann 1992)
P09955
- Class: Alpha Beta
- Architecture: 3-Layer(aba) Sandwich
- Topology: Aminopeptidase
- 34.3 kDa (Folk et al. 1960)
- 34.7 kDa (Theoretical)
9.0 (Wolff et al. 1962)
4.6-5.2 (Lipperheide and Otto 1986)
- 77,080-1M-1 (Theoretical)
- E1%,280 = 21.4 (Folk 1971)
- Glutamic acid (E378)
- Tyrosine (Y356)
- Competitively inhibited by arginine, lysine, and ornithine (Wolff et al. 1962)
- Metal chelating agents such as 1,10-phenanthroline and EDTA
- Heavy metals
- Not inhibited by diisopropylfluorophosphate (DFP) or phenylmethylsulfonyl fluoride (PMSF)
- Potato carboxypeptidase inhibitor (Hass et al. 1981)
- Leech carboxypeptidase inhibitor (Reverter et al. 1998)