For our international customers, please be advised that orders cannot be placed through our website by customers in countries with International Distributor representation.
Carboxypeptidase Y - Manual
Carboxypeptidase Y (CPDY) is a glycoprotein exopeptidase of the acid and serine class.
CPDY was originally referred to as proteinase C by Hata et al. in 1967. In 1970, Hayashi et al. found that CPDY released C-terminal amino acids from peptides and determined it is inactivated by DFP at the active site serine residue, thus classifying it as a serine peptidase (Hayashi et al. 1970, and Mortensen et al. 2004).
Wolf and Fink (1975) first identified the structural gene for CPDY, which was later confirmed by Hemmings et al. (1981). In the 1980s, extensive work was done to better understand the signal sequence of CPDY and it was determined that the protein can be translocated without its amino-terminal sequence (Hemmings et al. 1981 and Blachly-Dyson and Stevens 1987). Throughout the 1990s, the carbohydrate contents of CPDY were studied (Ballou et al. 1990), and in 1994 the crystal structure was solved to a 2.8 Å resolution by Endrizzi et al..
Recent work has focused on transport of the nascent protein (Gharakhanian et al. 2011, Mukaiyama et al. 2010, and Burston et al. 2008) and inhibitory proteins (Gombault et al. 2009).
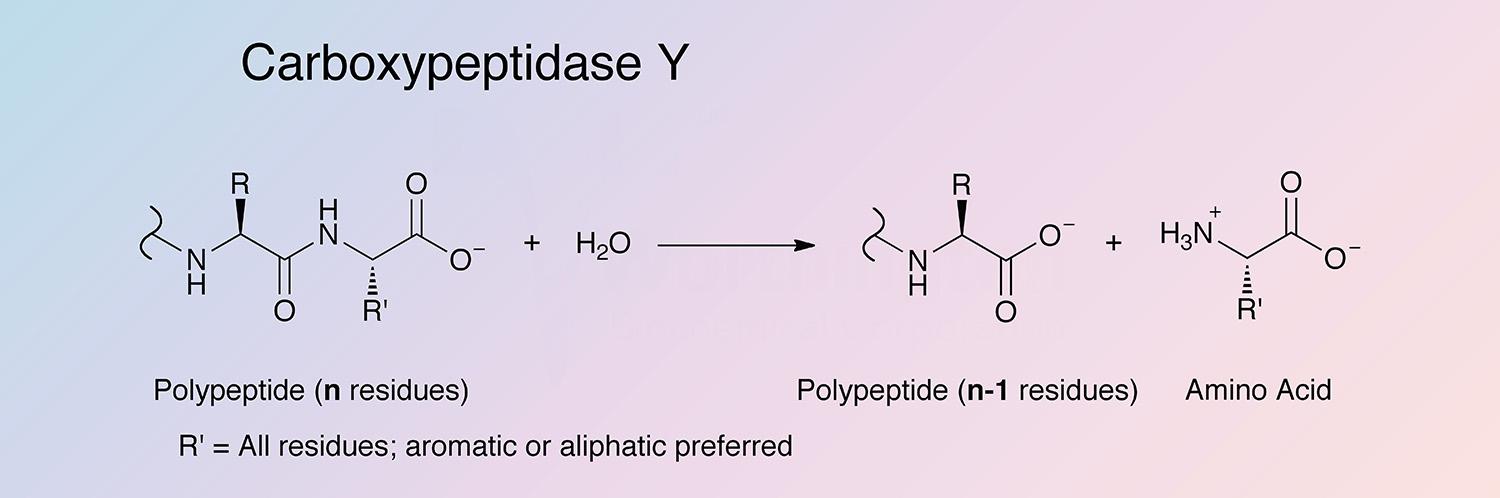
CPDY has broad amino acid specificity. It shows preference for hydrophobic amino acids in the P1’ position of the substrate. CPDY is also able to catalyze aminolysis, the reverse reaction of hydrolysis (Remington and Breddam 1994). For details on the random order bi-bi mechanism CPDY employs, see Mortensen et al. 1994.
Unlike carboxypeptidases A and B, CPDY contains no metal ion. It is glycosylated at four positions, and contains 15% mannose. Some of the carbohydrate chains are phosphorylated (Trimble and Maley 1977, Hasilik and Tanner 1978, Hashimoto et al. 1981, Winther et al. 1991, and Mortensen et al. 2004). Each molecule of CPDY contains 4-5 diesterfied phosphates (Hashimoto et al. 1981).
The CPDY structure consists of fourteen a-helices, eleven strands of mixed ß-sheets, five disulfide bridges, and one free cysteine residue (Endrizziet al. 1994).
The mature chain of CPDY contains 421 amino acid residues. The gene, prc1, has been cloned and sequenced and encodes a prepro form of the enzyme (Stevens et al. 1986, Blachly-Dyson and Stevens 1987, Valls et al. 1987, and Mortensen et al. 2004).
- C-terminal sequencing (Hayashi 1977)
- C-terminal modification/labeling of peptides and proteins (Remington and Breddam 1994)
P00729
CPDY contains two domains:
- Class: Alpha Beta; Mainly Alpha
- Architecture: 3-Layer(aba) Sandwich; Orthogonal Bundle
- Topology: Rossmann Fold; Helix Hairpins
- 64 kDa (Hayashi et al. 1973, and Mortensen et al. 2004)
4.5-6.0
3.6 (Hayashi et al. 1973)
- 88,940 cm-1M-1 (Theoretical)
- E1%,280 = 15.0 (Hayashi et al. 1973, and Kuhn et al. 1973)
- Serine (S146)
- Aspartate (D338)
- Histidine (H397)
- Diisopropyl fluorophosphate (DFP)
- PMSF
- APCK
- 4-Hydroxymercuribenzoate
- Aprotinin
- Sensitive to thiol-blocking reagents
- A high affinity inhibitor encoded by the TFSI gene has been characterized and shows homology to a family of lipid binding proteins (Bruun et al. 1998)