For our international customers, please be advised that orders cannot be placed through our website by customers in countries with International Distributor representation.
Glycerol Dehydrogenase - Manual
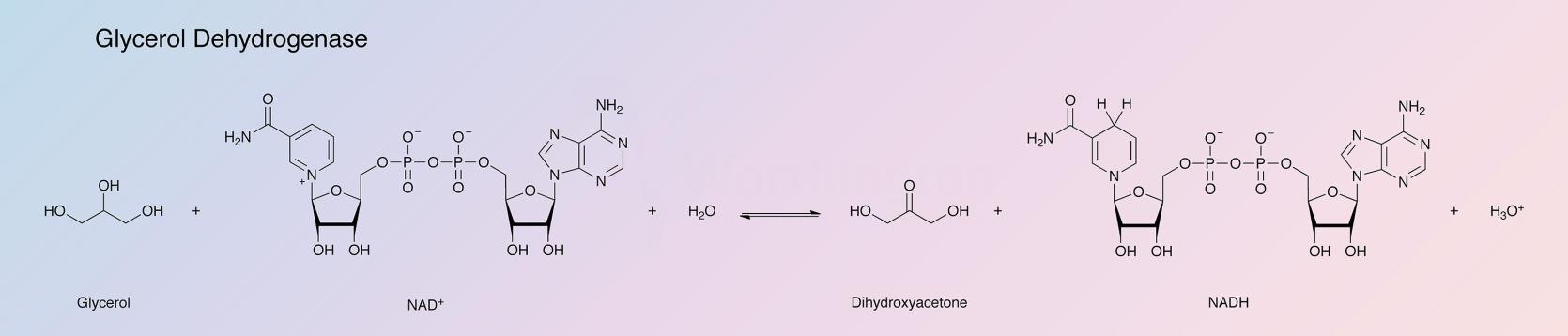
Glycerol dehydrogenase catalyzes the following reaction:
The primary interest in this enzyme is for the determination of glycerol in biological fluids: reduction of NAD is measured by change in absorbance at 340 nm.
Characteristics of Glycerol Dehydrogenase from Enterobacter aerogenes:
Method: The activity is measured using the procedure of Lin and Magasanik (1960) by determining the increase in absorbance at 340 nm resulting from the reduction of NAD. One unit reduces one micromole of NAD per minute at 25°C and pH 10.0 under the specified conditions.
Reagents
- 1.0 M Glycerol
- 0.125 M Potassium carbonate
- 0.125 M Sodium bicarbonate
- 0.125 M Carbonate/bicarbonate buffer, pH 10.0. Prepare by mixing potassium carbonate and sodium bicarbonate to reach pH 10.0.
- 1.0 M Ammonium sulfate
- 0.05 M Potassium phosphate, pH 7.6, containing 0.1 M ammonium sulfate and 0.1 mM manganese chloride
- 0.1 M NAD. Note: NAD may vary in salt form and degree of hydration. Care should be exercised to use an analytical grade and the correct molecular weight.
Enzyme
Dissolve enzyme at a concentration of one mg/ml in 0.05 M potassium phosphate buffer solution. Immediately prior to use dilute further in this buffer to obtain a rate of 0.02 - 0.05 ΔA/min.
Procedure
Adjust spectrophotometer to 340 nm and 25°C.
Pipette into each cuvette as follows:
1.0 M Ammonium sulfate 0.1 ml 0.1 M NAD 0.1 ml 0.125 M Carbonate/bicarbonate buffer, pH 10.0 2.4 ml 1.0 M Glycerol 0.3 ml
Incubate in the spectrophotometer for 4 - 5 minutes to achieve temperature equilibration and establish blank rate, if any. At zero time, add 0.1 ml of appropriately diluted enzyme and mix thoroughly. Record increase in A340 for 3 - 5 minutes. Determine ΔA340/min from initial linear portion of the curve.
Calculation
The greatest oxidizing activity is on glycerol and 1,2-propandiol. A table of substrate specificity is found in Lin and Magasanik (1960).
9.0 (Lin and Magasanik 1960). At pH 11, the enzyme is nearly inactive.
The enzyme is inhibited by zinc, (Lin et al. 1960); Li+ and Na+ and also by high ionic strength solutions (Strickland and Miller 1968).
The Worthington preparation is stable 6 - 12 months at 2 - 8°C. Concentrated solutions (10 mg/ml) are also relatively stable; on dilution to 1 mg/ml the activity rapidly disappears.
Assay
The Km for glycerol was found by Lin and Magasanik (1960) to be 0.017 M and 0.0056 M in the presence of 0.0033 M and 0.033 M ammonium chloride repectively. Vmax is increased by NH4+ also. They report that K+ and Rb+ activate the enzyme.