For our international customers, please be advised that orders cannot be placed through our website by customers in countries with International Distributor representation.
Deoxyribonuclease II - Manual
Porcine DNase II (pDNase II) is an acid endonuclease widely found in animal cells, mostly associated with lysosomes. It hydrolyzes the phosphodiester bonds of DNA under acidic conditions without the requirement for divalent metal ions and is important for DNA fragmentation and degradation.
In the late 1940s Catchside and Holmes first observed a mammalian DNase that was optimally active at an acidic pH (pH 4.5-5.5) and termed it “acid DNase” (Evans and Aguilera 2003).
Spleen tissue displays particularly high activity, so it was often referred to as “spleen acid DNase”. However, in 1968 Cordonnier and Bernardi found that acid DNase was found to be widely distributed in animal cells, and it was soon suggested that the enzyme be called DNase II to better distinguish it from pancreatic DNase (DNase I) (Cunningham and Laskowski 1953, and Evans and Aguilera 2003).
In the 1970s, Bernardi extensively investigated the inhibitory effects of various compounds on DNase II, finding actinomycin D, N-bromosuccinimide, and divalent cations to be inhibitory (Bernardi 1971). In 1973, Oshima and Price predicted the involvement of a histidine side chain in DNase II catalysis, and in 1985 Liao demonstrated this to be true.
In 1998, Liao first identified the structure of DNase II as a heterodimer, however it was soon shown by Wang et al. to be a heterotrimer (Liao et al. 1998, Wang et al. 1998). Yasuda et al. first characterized the human DNase II gene, the first DNase gene to be identified (Yasuda et al. 1998). Soon after, Baker et al. reported on cloning of the human gene as well (Baker et al. 1998).
Recent research has aimed at better understanding the role of DNase II in apoptosis (Kitahara et al. 2010) and its relevance to autoimmunity (Ueki et al. 2010).
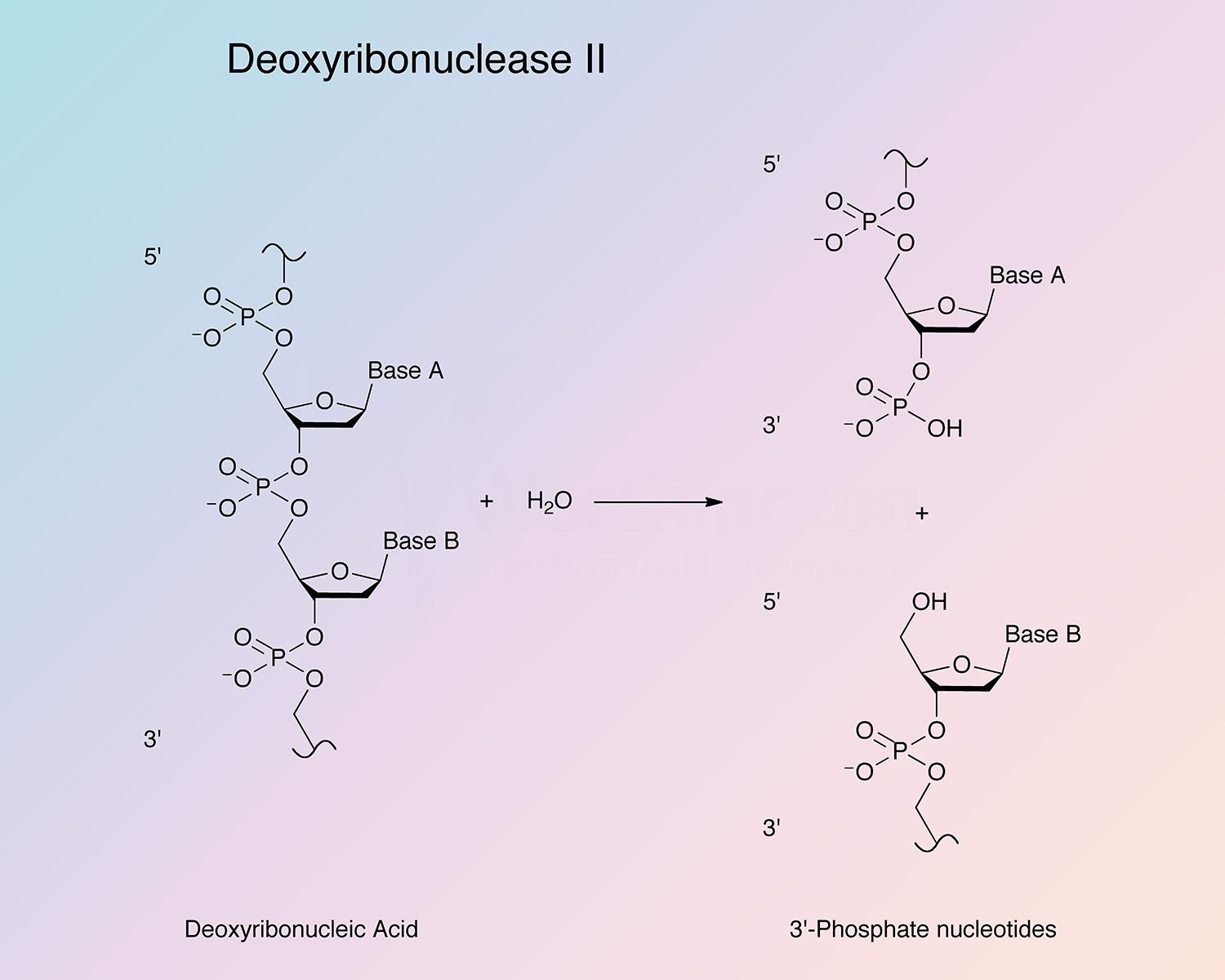
Through a nicking mechanism, DNase II generates 3’-phosphate groups by hydrolyzing the phosphodiester linkanges of DNA (Bernardi 1971, Harosh et al. 1991, and Baker et al. 1998).
A catalytic center, with a critical histidine residue, mediates the cleaveage (Oshima and Price 1973). Specifically, site-directed mutagenesis studies of histidine residues in pDNase II have suggested His115, His132, and His297 play crucial role in the enzyme’s function, and show absolute evolutionary conservation (Cheng et al. 2006).
The mechanism of DNase II has been shown to occur in three phases. Initially, multiple single strand breaks are induced within the DNA backbone. During the middle phase, acid soluble nucleotides and oligonucletides are produced. A slow, non-linear hyperchromic shift occurs in the last phase (Bernardi 1971).
It was initially discovered that the mature DNase II enzyme was a non-covalently bound alpha/beta heterodimer. It was proposed that during the production of endogenous pDNase II, the initial 45 kDa translation product is targeted to the lysosome where proteases cleave the translation product to two fragments of 35 kDa and 10 kDa, forming the dimer (Liao et al. 1989). However, Wang et al. showed that the two polypeptides of the alpha subunit dissociate into alpha1 and alpha2 subunits after disulfide reduction, forming a heterotrimeric structure composed of alpha1, beta, and alpha2 subunits (Wang et al. 1998, and Huang et al. 2009).
The genes encoding DNase II proteins have been described by Baker et al. 1998, Krieser and Eastman 1998, Shiokawa and Tanuma 1998, and Yasuda et al. 1998). The DNase2 gene is conserved in human, chimpanzee, dog, cow, mouse, rat, zebrafish, fruit fly, mosquito, and C. elegans. Porcine and human DNase II have been shown to share 73.4% identity (Shiokawa and Tanuma 1998).
The cDNA of pDNase II contains 1292 bases, with an open reading frame that encodes 364 amino acids, including six N-glycosylation sites (Shiokawa and Tanuma 1998).
To form the mature enzyme, a putative transmembrane peptide at the N-terminus is removed during the translation of the nascent protein. After protein folding, further processing within the lysosome removes a peptide at the C-terminus and two connecting peptides between the alpha1 and beta subunits and between beta and alpha2 (Wang et al. 1998). The resulting protein is a heterotrimer, as described in the Composition section.
- DNA fragmentation
- Apoptosis research
- Molecular weight marker
- Control in antibody reactions
O62855
- 38.0 kDa (Bernardi et al. 1965)
- 5.0 (although activity can be detected over a significant pH range) (Lyon and Aguilera 1997, Lyon et al. 2000)
- 4.5-5.0 at ionic strength 0.15 (Bernardi and Griffe 1964)
- 8.10 (Theoretical)
- 10.2 (Bernardi et al. 1965)
- 82,750 cm-1 M-1 (Theoretical)
- E1%, 280 = 12.1 (Bernardi et al. 1965)
- Histidine (H115, H132, H297) (Cheng et al. 2006)
- Cysteine (Bernardi 1971)
- EDTA (Bernardi 1971)
- Actinomycin D (Bernardi 1971)
- Iodoacetate (Liao et al. 1989)
- N-Bromosuccinimide and H2O2 (Bernardi 1971)
- Divalent cations such as zinc and copper (Bernardi 1971)
- Sulfate (Bernardi and Griffe 1964)
- NaCl (above 250 mM) (Ikeda 2002)
- DNA (above .4 mg/ml), rRNA, tRNA (Bernardi 1971)
- Iodoacetamide (slightly inhibitory) (Bernardi 1971)