For our international customers, please be advised that orders cannot be placed through our website by customers in countries with International Distributor representation.
Reverse Transcriptase, Recombinant HIV - Manual
Reverse transcriptases are enzymes encoded in retroviruses viral genome. The enzyme is responsible for transcription of the viral RNA to produce a dsDNA that can be inserted into the host genome.
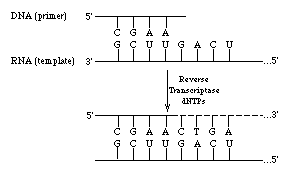
Reverse transcriptases are multifunctional enzymes. These enzymes exhibit an RNA and DNA directed polymerase activity. In addition reverse transcriptases catalyze the degradation of RNA in an RNA-DNA hybrid. The exonucleolytic activity proceeds in a 5' ---> 3' direction. The RNA or DNA directed activity requires a template (RNA or DNA) and a primer. The following is a schematic illustration of the reaction:
Characteristics of Reverse Transcriptase from recombinant E. coli
The recombinant HIV reverse transcriptase consists of a heterodimer , with subunits having a MW of 66,000 and 51,000 daltons (p66/p51 heterodimer)
Optimal activity is obtained at pH 8. Mg++ ions and sulfhydryl reagents are required. For RNA transcription a RNA template and deoxyoligonucleotides primer are required. The primer may be oligo dT 8-12 or a mixture of DNAs with random or specific sequences. The resulting DNA transcribed is referred to as complementary DNA (cDNA). Reverse transcriptases exhibit a DNA directed DNA polymerase. This activity lacks the 3' ---> 5' exonuclease activity usually associated with bacterial DNA dependent DNA polymerase.
A third activity of the HIV reverse transcriptase is RNase H. RNase H degrades RNA in RNA:DNA hybrid. Analysis of the structure-function relqationship between the polymerase and RNase H activities indicates that the two activities reside in separate segments of the molecule. While the polymerase domain is contained within the NH2-terminal 51KDa, the RNase H activity is located in COOH-terminal 15 KDa of the p66 subunit.
HIV reverse transcriptase is used for research on the AIDS primer. However it can be substituted for AMV reverse transcriptase, which is mainly used to transcribe mRNA into double stranded cDNA, that can be inserted into prokaryotic vectors. The enzyme can also be used with either single stranded DNA or RNA templates to make probes for use in hybridization experiments. It can be used for labeling the termini of DNA fragments with protruding 5' termini. The enzyme can also be used to sequence DNAs by the dideoxy chain termination method of Sanger when the Klenow fragment of E. coli DNA polymerase I, or the T7 DNA polymerase yield unsatisfactory results.