For our international customers, please be advised that orders cannot be placed through our website by customers in countries with International Distributor representation.
Pepsin - Manual
Pepsin is the principal proteolytic enzyme of vertebrate gastric juice. Its inactive precursor form, pepsinogen, is produced in stomach mucosa. The minor pepsins are designated “B”, “C”, and “D”, while the major component is “A”, to which the following data applies.
Pepsin is of particular interest as it was the first enzyme to be discovered. The name pepsin was given by Theodor Schwann (1810-1882) in 1836, and came from pepsis, the term for digestion in Hippocratic writings. Into the mid-nineteenth century, scientists showed that pepsin broke down proteins into “peptones” (Fruton 2002).
Pepsin was later found to be an effective treatment for digestive disorders. Through this important application, efforts to produce and purify it greatly increased, and were successful by the end of the nineteenth century (Tang 1998).
At that time, however, the chemical nature and properties of enzymes as proteins were not completely understood. It was not until John H. Northrop crystallized pepsin in 1930, an achievement for which he shared the Nobel Prize in 1946, that the protein nature of enzymes was established (Manchester 2004).
After the Nobel Prize was awarded to Northrop, Sumner, and Stanley in 1946, new separation methods including crystallization and chromatography were further developed. Through these methods, the amino acid sequences of pepsin and pepsinogen were determined (Tang 1973).
Pepsin B and C were first isolated from porcine stomach by Ryle and Porter in 1959.
As X-ray diffraction techniques improved through the mid-1970s, the three-dimensional structure of pepsin was determined, allowing for a better understanding of the catalytic reaction (Fruton 2002).
Recently, interest in pepsin-type enzymes and their inhibitors has been renewed due to the recognition of HIV-protease as a member of this aspartic protease family (Campos 2003).
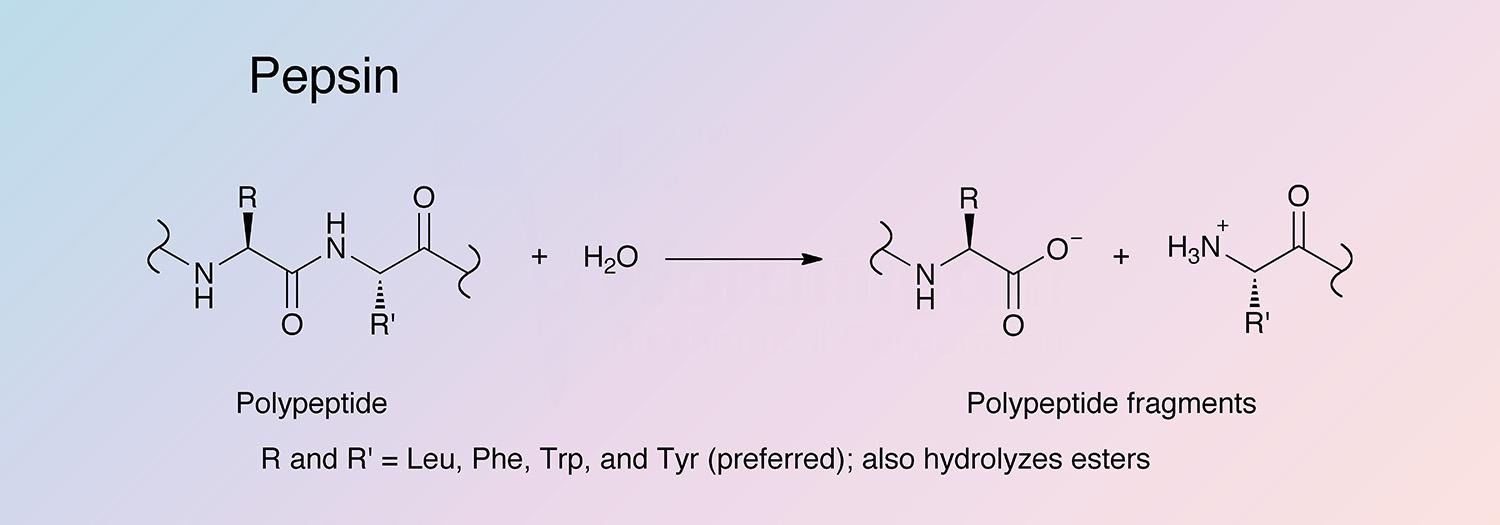
Pepsin has broad specificity with a preference for peptides containing linkages with aromatic or carboxylic L-amino acids. It preferentially cleaves C-terminal to Phe and Leu and to a lesser extent Glu linkages. The enzyme does not cleave at Val, Ala, or Gly.
Molecular Characteristics:
The amino acid sequence of porcine pepsin was determined by Tang et al. (1973) and Moravek and Kostka (1974), and later confirmed through cDNA analysis by Tsukagoshi et al. (1988) and Lin et al. (1989).
The pepsinogen A (PGA) gene is divided among nine exons that encompass approximately 9.4 kb of genomic DNA (Sogawa 1983).
There are multiple versions of the PGA genes found in human and chimp populations, but the activities of these various gene products are indistinguishable (Taggart 1985 and Zelle 1988). In contrast, Southern blot analyses of a sampling of pigs suggest that there is only a single PGAgene found in all pigs (Evers 1988).
PGA production is mainly controlled at the transcription level (Sogawa et al. 1981 and Ichinose et al. 1988). In both humans and pigs, it has been found that the PGA gene is under tissue-specific transcriptional control, with mRNA only detected in gastric fundic mucosa (Ichinose 1991 and Meijerink et al. 1993). Transcription of the PGA gene is regulated by transcription-activating proteins acting at 3 major regions in the promoter and initiation regions of the PGA gene (Meijerink et al. 1993).
There are four reported pepsin proteins: pepsin A, pepsin B (parapepsin I), pepsin C (gastricsin), and pepsin D (an unphosphorylated version of pepsin A) (Lee and Ryle 1967). Pepsin A is the predominant gastric protease; minor amounts of the other pepsins have been detected. Pepsins B and C share a higher degree of homology with each other. In dog, B and C share 89% identity, A and B share 44% identity, and A and C share 45% identity (calculated based on Thompson et al. 1994).
Pepsin is a monomeric, two domain, mainly beta protein with a high percentage of acidic residues. Porcine pepsin has 4 basic residues, and 42 acidic residues and is O-phosphorylated at S68 (Tang et al. 1973). For the protein to be active, one of the two aspartate residues in the catalytic site has to be protonated, and the other deprotonated. This occurs between pH 1 and 5, and above pH 7 pepsin is irreversibly denatured.
- Digestion of antibodies
- Preparation of collagen for cosmeceutical purposes
- Assessment of digestibility of proteins in food chemistry
- Subculture of viable mammary epithelial cells (Riser 1983)
P00791
- Class: Mainly beta
- Architecture: Beta Barrel
- Topology: Cathepsin D, subunit A; domain 1
- Pepsin: 34.5 kDa (Theoretical)
- Pepsinogen: 41.4 kDa
1.0-4.0 (At pH 1.5 pepsin exhibits about 90% of maximum activity, and at pH 4.5 about 35% of maximum activity.
1.0 (Bovey and Yanari 1960)
- 49,650 cm-1 M-1 (Theoretical)
- E1%,280 = 14.39 (Theoretical)
- Aspartic acid (D32 and D215)
- Pepsinogen
- Aliphatic alcohols
- Substrate-like epoxides
- Pepstatin A