For our international customers, please be advised that orders cannot be placed through our website by customers in countries with International Distributor representation.
Ribonuclease T1 - Manual
Ribonuclease T1 is a low molecular weight, highly specific, monomeric endoribonuclease. That from the fungus A. oryzae is the leading member of the family of microbial enzymes that cleave single stranded RNA by a reaction mechanism involving formation of intermediate 2’,3’ cyclic phosphates.
In 1957, Sato and Egami first discovered ribonuclease T1 in a commercial enzyme mixture from A. oryzae called Takadiastase. The primary structure was determined (Takahashi 1971a, b) and revised and confirmed in 1985 (Takahashi 1985).
In the late 1980s, the enzyme was well characterized by NMR spectroscopy, chemical modification, and x-ray crystallography (Hoffman and Rüterjans 1988, Arnie et al. 1988, Koepke et al. 1989). Although the gene encoding RNase T1 had not yet been cloned, DNA corresponding to the full amino acid sequence of mature RNase T1 was synthesized and expressed in E. coli as a fusion protein (Ikehara et al. 1986, and Quaas et al. 1988). Using chemically synthesized DNA, researchers also modified the active sites to investigate the relationship between structure and function (Nishikawa et al. 1986, and Hakoshima et al. 1988). The gene (rntA) was cloned and sequenced by Fuji et al. in 1995.
RNase T1 continues to be used in RNA mapping experiments, aiding in the determination of RNA secondary structure (Quarrier et al. 2010) and identification of a given RNA molecule’s origin (Matthiesen and Kirpekar 2009). RNase T1 has been used as a model protein for structural studies (Moors et al. 2009, and Yoshida et al. 2006).
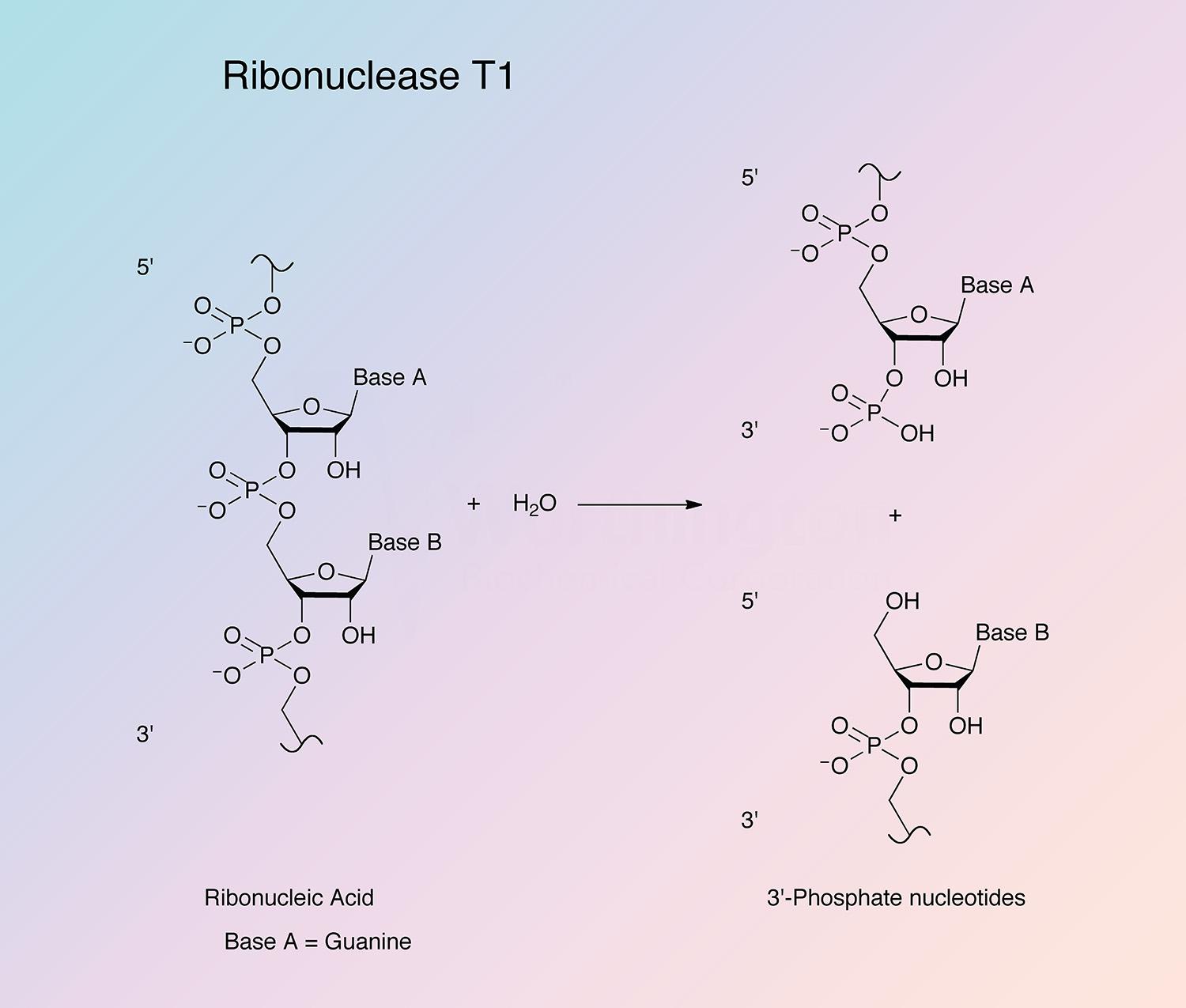
RNase T1 is highly specific in its mode of attack. The enzyme first cleaves RNA (or deaminated RNA) between 3’-guanylic residues (or inosine 3’-phosphate and xanthosine 3’-phosphate) and the 5’-hydroxy residues of adjacent nucleotides with the formation of the corresponding intermediate 2’,3’-cyclic phosphates (Takahashi and Moore 1971). This step is reversible and is much faster than the second step. In the second step, the intermediate is hydrolyzed to a 3’-phosphate (Yoshida 2001). All members of the RNase T1 family are guanyl specific or preferential, except RNase U2. It is believed that the highly conserved region from Tyr-42 to Glu-46 is primarily responsible for guanine binding (Yoshida 2001).
RNase T1 is a low molecular weight, stable, monomeric protein. The architecture consists of four antiparallel beta strands over an alpha helix (Yoshida 2001).
RNase T1 is composed of 104 amino acid residues cross-linked by two disulfide bridges. A conserved disulfide bond exists between Cys-6 and Cys-103 (Yoshida 2001). The sequences of RNase N1 (from Neurospora crassa), and U1 (from Ustilago sphaerogena) are highly homologous to RNase T1 (Takahashi 1985). The nucleotide sequence of the rntA gene contains an open reading frame of 393 bp with one intervening intron (114 bp). The intron is present within the region corresponding to the signal sequence. A sequence related to the TATA box (TATAA) is present at nucleotides 121 to 117, and a CAAT box is present at nucleotides 176 to 173 (Fujii et al. 1995).
- Cleavage enzyme for the determination of structure, nearest neighbor frequencies, and sequence of RNA (Miura and Egami 1960, Rushizky and Sober 1962, Miura 1964b, Holley et al. 1965, Madison and Kung 1967, and Neelon et al. 1967)
- Preparation of nucleoside 2’,3’-cyclic phosphates
- Synthesis of oligonucleotides
- Removal of RNA from DNA preparations (Saito and Miura 1963)
- Removal of poly(A) sequences from mRNA in the presence of oligo(dT)
- Removal of mRNA during single-strand cDNA synthesis
- RNA fingerprinting
- Oligodeoxyribonucleotide-directed cleavage of RNA
P00651
- Class: Alpha Beta
- Architecture: Roll
- Topology: Nuclear Transport Factor 2; Chain A
- 11 kDa (Egami et al. 1964)
- 11.1 kDa (Theoretical)
7.5
- 2.9 (Egami et al. 1964)
- 4.12 (Theoretical)
- 17,690 cm-1 M-1
- E1%,278 = 2.9 (Egami et al. 1964)
- Histidine (H40 and H92)
- Glutamic acid (E58)
- Arginine (R77)
- Histidine
- EDTA
- Ag+, Zn2+, Cu2+, and Hg2+ at 1 x 10-3 M (strongly inhibitory)