For our international customers, please be advised that orders cannot be placed through our website by customers in countries with International Distributor representation.
Alcohol Dehydrogenase - Manual
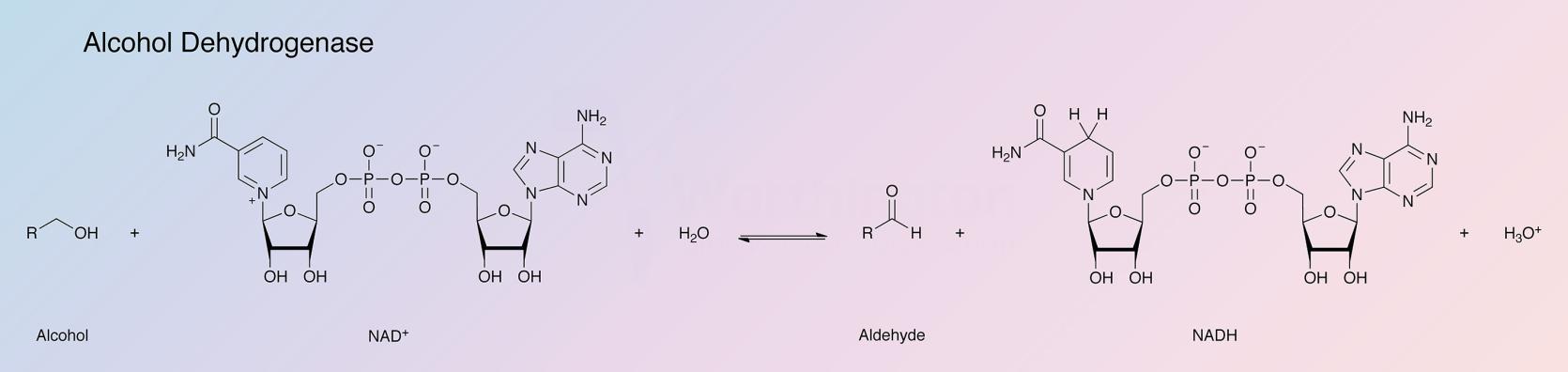
Alcohol dehydrogenase (ADH), part of the oxidoreductase family, catalyzes the oxidation of alcohols, using NAD+ or NADP+ as the electron acceptor (White and White 1997). The reaction is reversible and substrates can be a variety of primary or secondary alcohols, and hemiacetals. Alcohol dehydrogenases are present in most organisms, with that of yeast being the most active form of the enzyme. The following data applies to the alcohol dehydrogenase of baker's yeast, Saccharomyces cerevisiae.
In 1937 alcohol dehydrogenase was first purified and crystallized from brewer's yeast by Negelein and Wulff (Negelein and Wulff 1937). In 1948, Bonnichsen and Wassen crystallized ADH from horse liver (Bonnichsen and Wassen 1948). It was found that these two ADHs differed in many of their properties.
In the early 1950s Theorell and Chance studied the stoichiometry and dissociation constant of the mammalian enzyme complex. It was found that the yeast enzyme is twice the size of mammalian ADH, and approximately 100 times more active (Hayes and Velick 1953). In 1955, Vallee and Hoch confirmed the presence of Zinc metal (Vallee and Hoch 1955a).
In the 1960s the inhibitors of alcohol dehydrogenase were studied (Atkinson et al. 1967), along with the roles of specific structural components (Auricchio and Bruni 1969). Structural and kinetic studies (Blackwell et al. 1974) continued on into the 1970s when conformational changes associated with binding were investigated (Abdallah et al. 1975), along with the isozymes (Berger et al. 1974).
During the 1980s the genetics, biochemistry, and developmental regulation of ADH in various species, including mice (Watabiki et al. 1989) and pig (Keung and Fong 1988), were investigated. The 1990s brought better understanding of the role of the zinc metal (Magonet et al. 1992), and the discovery of additional inhibitors (Pereira et al. 1992, Sachan and Cha 1994, and Shiao et al. 1994).
Recent research has focused on obtaining alcohol dehydrogenases with higher catalytic activity, and a better understanding ADH gene regulation (Larroy et al. 2002a and b, Lertwattanasakul et al. 2007, and Park 2009).
Yeast ADH has a more narrow specificity than that of the liver enzyme. It accepts ethanol and is somewhat active on the straight chain primary alcohols. It acts to a very limited extent on certain secondary and branched chained alcohols (Dickinson and Dalziel 1967a).
Alcohol dehydrogenase is a tetramer with each subunit containing one zinc atom (Vallee and Hoch 1955). Per subunit, there are two distinct active site sulfhydryl groups which can be distinguished on the basis of differential reactivity with iodoacetate and butyl isocyanate (Twu, Chin, and Wold 1973). A histidine residue has an essential role (Dickenson and Dickinson 1975 and LeBrun et al. 2004).
Seven genes for Saccharomyces cerevisiae have been identified (Lertwattanasakul et al. 2007). ScADH1 is expressed in large amounts in the presence of glucose, and it encodes for the fermentative enzyme that produces ethanol (Bennetzen and Hall 1982). ScADH2 is negatively regulated by glucose and encodes the isozyme that converts ethanol to acetaldehyde (Ciriacy 1975, Wills and Jornvall 1979, and Russel and Hall 1983). The ScADH3 gene is suppressed by glucose and the mature form of the protein is found the mitochondria (Young and Pilgrim, 1985). ScADH4, ScADH5, and ScADH6 encode for the ScADH4, ScADH5, and ScADH6 proteins. ScADH6 and ScADH7 are thought to contribute to the balance of NADP/NADPH (Larroy et al. 2002a and b).
- Enzymatic determination of primary alcohols, and aldehydes
- Synthesis of chiral compounds
- Spectrophotometric assay of plasmalogenase
- Enzymatic catalysis in organic solvents
- Studies of NAD+, NADH, NADP+, and NADPH
P00330
- Class: Alpha Beta
- Architecture: Alpha-Beta Complex and 3-Layer (aba) Sandwich
- Topology: Quinone Oxidoreductase; Chain A, domain 1 and Rossmann fold
146.8 kDa (Theoretical)
5.4 (Shore and Theorell 1966)
6.23 (Theoretical)
- 189,320 cm-1 M-1 (Theoretical)
- E1%,280 = 12.89 (Theoretical)
- Sulfhydryl activating reagents
- Mercaptoethanol
- Dithiothreitol
- Cysteine
- Heavy metal chelating agents
(White and White 1997)
- Heavy metals and -SH reagents
- Thiourea
- Purine and pyrimidine derivatives
- Chloroethanol and fluoroethanol
- N-alkylmaleimides
- Iodoacetamide
- 1,10-phenanthroline
- 8-hydroxyquinoline
- Beta-NAD analogs
(White and White 1997)