For our international customers, please be advised that orders cannot be placed through our website by customers in countries with International Distributor representation.
Papain - Manual
Papain is a cysteine hydrolase that is stable and active under a wide range of conditions. It is very stable even at elevated temperatures (Cohen et al. 1986). The latex of Carica papaya is a rich source of four cysteine endopeptidases including papain, chymopapain, glycyl endopeptidase, and caricain. The proteins are synthesized as inactive precursors that become active within two minutes of the plant being wounded and the latex expelled. Papain is a minor constituent, but has been more widely studied because it is more easily purified (Azarkan 2003).
In 1873 G.C. Roy first investigated the action of papain in an article published in the Calcutta Medical Journal entitled “The Solvent Action of Papaya Juice on Nitrogenous Articles of Food”. Papain was first named in the late nineteenth century by Wurtz and Bouchut who partially purified the product from the sap of papaya (Menard and Storer 1998). When named, it was simply recognized as a proteolytically active constituent in the latex of tropical papaya fruit (Wurtz and Bouchut 1879).
Throughout the mid-1950s and 1960s, purification and separation techniques improved greatly and pure papain was isolated. The study of papain allowed for great advances in understanding enzymes as proteins. In 1968, papain was the second enzyme to be crystallized and its structure determined by x-ray methods. Papain was the first cysteine protease to have its structure identified (Drenth et al. 1968).
In the 1980s, the geometry of the active site was reviewed and the three-dimensional structure was determined to a 1.65 Angstrom resolution (Kamphuis et al. 1984). The precursors and inhibitors of papain were extensively studied into the 1990s (Vernet et al. 1991).
Today’s research aims to further understand the specificity (Portaro et al. 2000) and the structural perturbations brought about by inhibitors, low pH, metal ions, and fluorinated alcohols (Alphey and Hunter 2006, Huet et al. 2006, Kaul et al. 2002, Naeem et al. 2004).
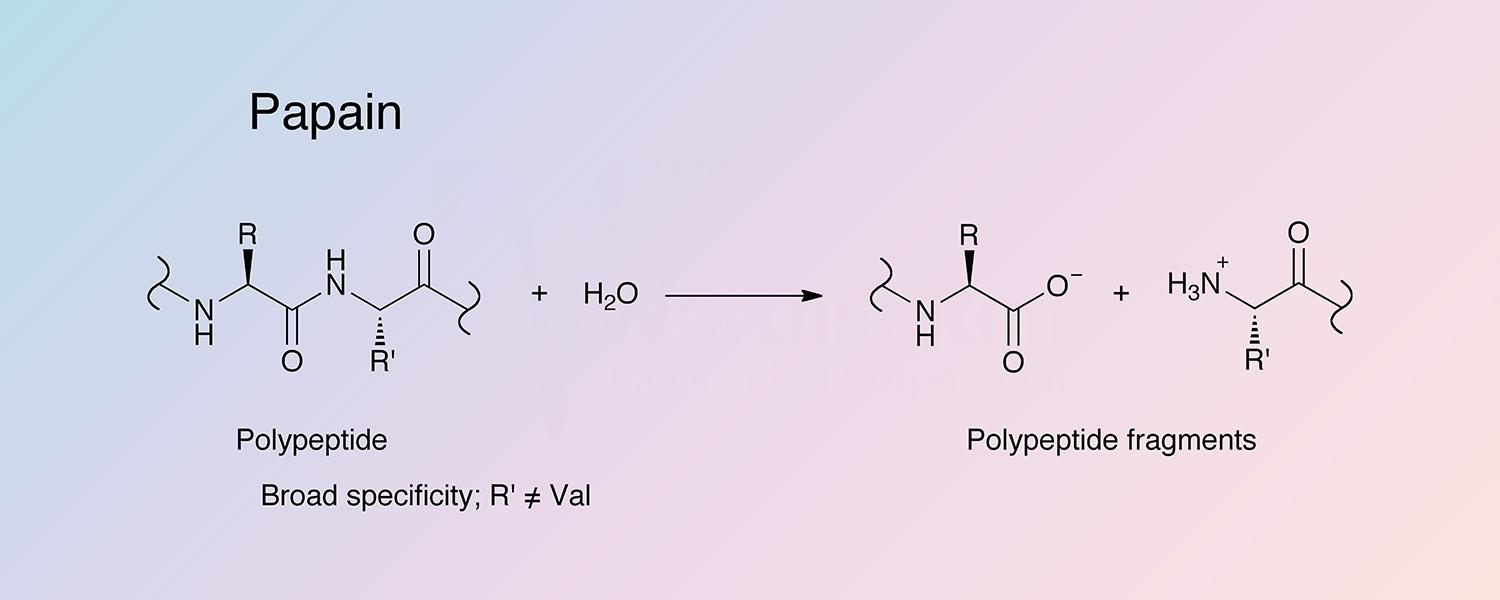
Papain has fairly broad specificity; it has endopeptidase, amidase, and esterase activities. The active site consists of seven subsites (S1-S4 and S1’-S3’) that can each accommodate one amino acid residue of a substrate (P1-P4 and P1’-P3’) (Schechter and Berger 1967). Specificity is controlled by the S2 subsite, a hydrophobic pocket that accommodates the P2 side chain of the substrate. Papain exhibits specific substrate preferences primarily for bulky hydrophobic or aromatic residues at this subsite (Kimmel and Smith 1954). Outside of the S2 subsite preferences, there is a lack of clearly defined residue selectivity within the active site.
The mature forms of all papaya proteinases are between 212 and 218 amino acids, and exhibit a strong degree of homology (Azarkan 2003). X-ray structure analysis has shown that they adopt identical three-dimensional folds (Pickersgill et al. 1991, O’Hara et al. 1995, and Maes et al.1996).
Papain is synthesized as a zymogen with a 133 amino acid N-terminal region that is not part of the active enzyme (Cohen et al. 1986). The papain precursor gene, prepropapain, has been cloned and expressed either in parts or as a whole (Cohen et al. 1986 and Choudhury et al.2009).
Composition:
Papain is a single-chained polypeptide with three disulfide bridges and a sulfhydryl group necessary for the activity of the enzyme. Papain is expressed as an inactive precursor, prepropapain. The formation of active papain requires several cleavage steps including an initial cleavage of the 18 amino acid preregion (the signal sequence), followed by further cleavage of the glycosylated 114 amino acid proregion (Vernet et al. 1995). This proregion serves as an intrinsic inhibitor and folding template. For further detail see Revell et al. 1993, Taylor et al. 1992, and Cohen et al. 1990.
- Cell isolation where it is more gentle than other proteases (i.e.: cortical neurons, retina, and smooth muscle)
- Protein structural studies, peptide mapping
- Red cell surface modification for antibody screening or identification
- Fab preparation from IgG and IgM antibodies
- Solubilization of integral membrane proteins
- Production of glycopeptides from purified proteoglycans
- Enzymatic wound debridement
P00784
- Class: Alpha Beta
- Architecture: Alpha-Beta Complex
- Topology: Cathepsin B; Chain A
- 23.4 kDa (Theoretical)
6.0-7.0
- 8.88 (Theoretical)
- 53,610 cm-1 M-1 (Theoretical)
- E1%,280 = 22.88 (Theoretical)
- Cysteine (C158)
- Histidine (H292)
- Asparagine (N308)
- Cysteine
- Sulfide and sulfite
- Heavy metal chelating agents like EDTA
- N-bromosuccinimide
(White and White 1997)
- PMSF
- TLCK, TPCK
- alph2-macroglobulin
- Hg2+ and other heavy metals
- AEBSF
- Antipain
- cystatin
- E-64
- Leupeptin
- Sulfhydryl binding agents
- Carbonyl reagents
- Alkylating agents
(White and White 1997)