For our international customers, please be advised that orders cannot be placed through our website by customers in countries with International Distributor representation.
Trypsin - Manual
Trypsin is a pancreatic serine protease with substrate specificity based upon positively charged lysine and arginine side chains (Brown and Wold 1973). The enzyme in excreted by the pancreas and takes part in the digestion of food proteins and other biological processes. Trypsin is a medium-sized globular protein and is produced as an inactive proenzyme, trypsinogen (Chen et al. 2009).
In 1876, trypsin was first named by Kuhne who described the proteolytic activity of this pancreatic enzyme. He compared trypsin and pepsin, discovering the differentiating factor to be the optimal pH. In 1931, Northrop and Kunitz purified trypsin by crystallization shortly after first purifying pepsin in 1930.
In 1974 the three dimensional structure was determined, which served as a prototype for the serine endopeptidase S1 family to which trypsin belongs.
In the late 1980s, and early 1990s, site directed mutagenesis with recombinant trypsin determined the role of particular amino acid residues (Sprang et al. 1987, McGrath et al. 1989, Corey and Craik 1992, and Corey et al. 1992).
In the late 1990s trypsin’s role in hereditary pancreatitis was investigated, and it was determined that a mutation at Arg117His is responsible for preventing autolysis thereby causing pancreatitis.
Today, trypsin continues to be used in the development of cell and tissue culture protocols (Soleimani et al. 2009, Banumathi et al. 2009, and Yang et al. 2009), as well as protein identification through peptide sequencing techniques (Manz et al. 2004 and Schuchert et al. 2009). In the medical field, the role of trypsin in pancreatic diseases, including cystic fibrosis (Tzetis et al. 2007, and Li et al. 2009) and chronic pancreatitis (Chen et al. 2009), has been the subject of current research, and trypsin has been used to model the decomposition of articular cartilage in osteoarthritis (Wang et al. 2008).
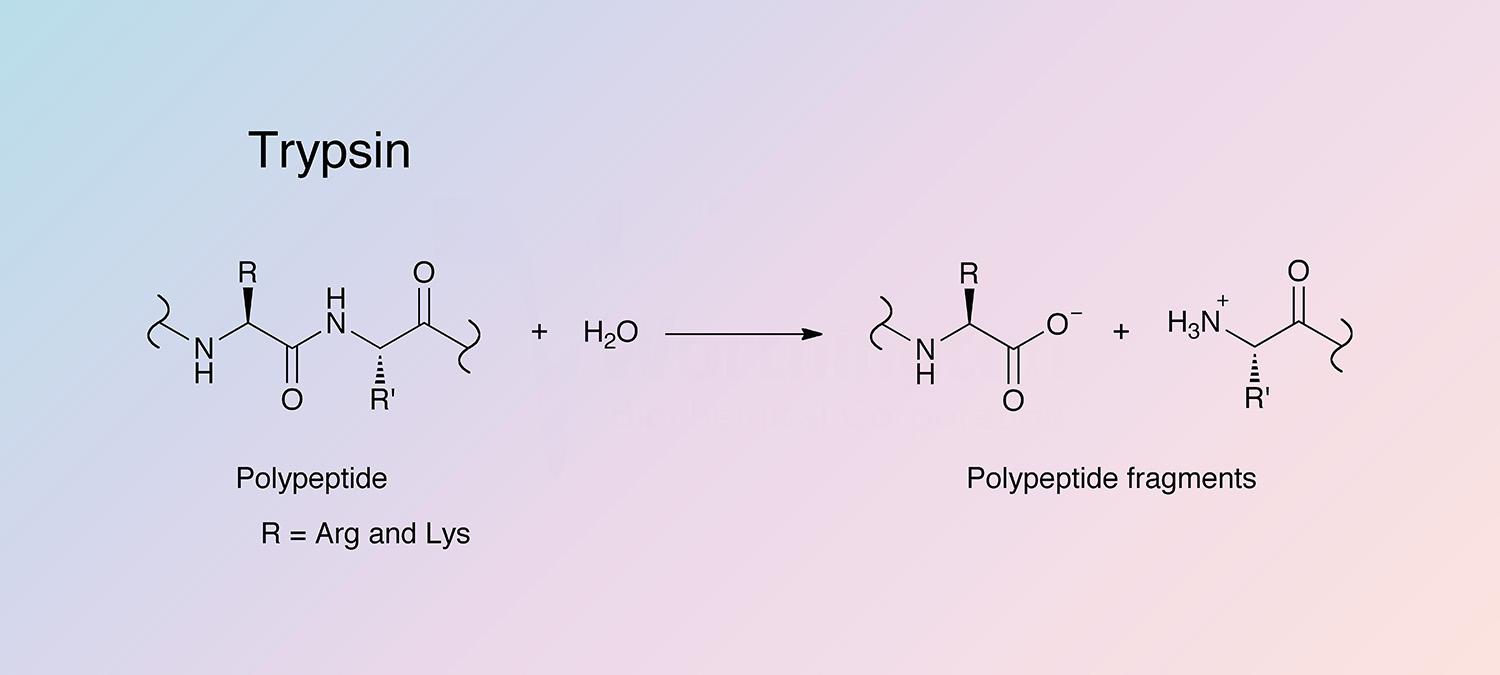
Trypsin cleaves peptides on the C-terminal side of lysine and arginine amino acid residues. If a proline residue is on the carboxyl side of the cleavage site, the cleavage will not occur. If an acidic residue is on either side of the cleavage site, the rate of hydrolysis has been shown to be slower.
Trypsinogen may be activated by removal of a terminal hexapeptide to yield single-chain β-trypsin. Subsequent limited autolysis produces other active forms having two or more peptide chains bound by disulfide bonds. The predominant forms are α-trypsin, having two peptide chains and β-, a single chain. Different activity and thermal stability are shown by α- and β-trypsin.
Other structural features include surface loops at amino acids 185-193, which influence specificity, despite not making direct contact with the substrate. A high affinity Ca2+ binding site is required for stability, and when not present, autolysis occurs. The autolysis loop (located at amino acids 143-151) is very flexible in both trypsin and trypsinogen. Cleavage at the lysine yields the alpha form which retains some catalytic activity. The protein has six completely conserved disulfide bonds (Halfon and Craik 1998).
Bovine pancreas expresses two forms of trypsin, the dominant cationic and minor anionic forms. These protein sequences share 72% identity, while their coding regions share 78% identity.
Each of these proteins are further processed into alternate forms. Catalytic trypsin contains a flexible “autolysis loop” (residues G145-V157) (Schroeder and Shaw 1968, and Bartunik et al. 1989), and autolysis of the dominant, single-chain form B-trypsin at K148-S149 within this loop leads to the formation of A-trypsin. Further autolysis at K193-D194 leads to the formation of Psi-trypsin (Fehlhammer and Bode 1975).
Both the cationic and anionic trypsin proteins are expressed as trypsinogen proenzymes, with a 15-residue signal peptide (M1-A15) and an 8-residue propeptide (F16-K23). The three-dimensional fold of all known trypsins is highly conserved. In addition, the catalytic triad and regions flanking the catalytic triad are highly conserved (Hartley 1970).
- Tissue dissociation, especially when combined with other enzymes such as collagenase, and elastase
- Cell harvesting by “trypsinization”
- Mitochondria isolation
- in vitro studies of proteins
- Removing monolayers of cells from plastic and glass
- Various hemagglutination procedures
- Sample preparation for flow cytometric DNA analysis
- Tryptic mapping
- Fingerprinting and sequencing work
- Environmental monitoring
- Reduction of cell density in tissue culture
- Subculturing cells
- Cleavage fusion proteins
- Generating glycopeptides from purified glycoproteins
(White and White 1997)
P00760
- Class: Mainly Beta
- Architecture: Beta Barrel
- Topology: Thrombin, subunit H
23.3 kDa (Theoretical)
7.5-8.5 (Koutsopoulos et al. 2007)
- Trypsinogen: pH 9.3 (Walsh and Neurath 1964)
- Trypsin: pH 10.5 (Cunningham 1954).
Trypsinogen: 45,250 cm-1 M-1>
- Histidine (H63)
- Aspartic acid (D107)
- Serine (S200)
The rate of trypsinogen conversion is enhanced by using lanthanide in place of calcium ions (Gomez et al. 1974).
- Pancreatic-, soybean-, lima bean-, and egg white- trypsin inhibitors (see section on Trypsin Inhibitors)
- DFP
- Aprotinin
- Ag+
- Benzamidine
- EDTA
(White and White 1997)